Demonstration of single- and multi-plex 6-log dynamic range with Countable PCR
Introduction
Countable PCR enables true single-molecule counting by spatially isolating individual target molecules within a 3D gel-like matrix inside a standard PCR tube. This matrix comprises over 30 million compartments, which gives key advantages for gene quantification:
- A six-log dynamic range that greatly surpasses the dynamic range of legacy digital PCR (dPCR) systems without requiring Poisson correction.
- Unbiased, enhanced, and scalable multiplexing across a six-log dynamic range, made possible by single-molecule occupancy.
Here, we demonstrate each of these principles on the Countable PCR system.
Materials and Methods
Comparing Countable PCR with legacy digital PCR instruments
Short (<200 bp) synthetic double-stranded DNA templates derived from RPP30 were prepared and diluted to counts between 100 and 1,000,000 molecules. PCR reactions were set up to quantify those dilutions using 1) Countable PCR, 2) a droplet dPCR system, and 3) a nanoplate dPCR system.
Each reaction used identical RPP30 TaqMan primers and probes as described in Table 1. Four replicates of each dilution were quantified on each system. Thermal cycling conditions for all systems are described in Table 2. The Countable PCR reaction was set up according to Table 3, and the reactions on the other dPCR instruments were set up according to the manufacturer’s recommended protocol.
Single-plex and four-plex performance on Countable PCR
Four short (<200 bp) synthetic double-stranded DNA templates, derived from four human genes (RPP30, JAK2, RAD51, and MET), were prepared and serially diluted to achieve counts ranging from 10 to 1,000,000 molecules. For the multiplex reactions, a pool of TaqMan assays for each target was prepared for Ch01, Ch02, Ch03, and Ch04, respectively (Table 1). The pool of synthetic templates, ranging from 10 to 1,000,000 molecules, was added to a series of multiplex PCR reactions containing the pool of TaqMan assays (Table 4). Eight replicates of each dilution were quantified with Countable PCR.
For the single-plex reactions, serially diluted JAK2 templates (as described above) were added to a single-plex reaction containing the JAK2 TaqMan assay for Ch02 (Table 5). Each dilution was quantified in eight replicates using Countable PCR.
Thermal cycling for all samples was performed as described in Table 2.
Table 1. Primers for dynamic range experiments
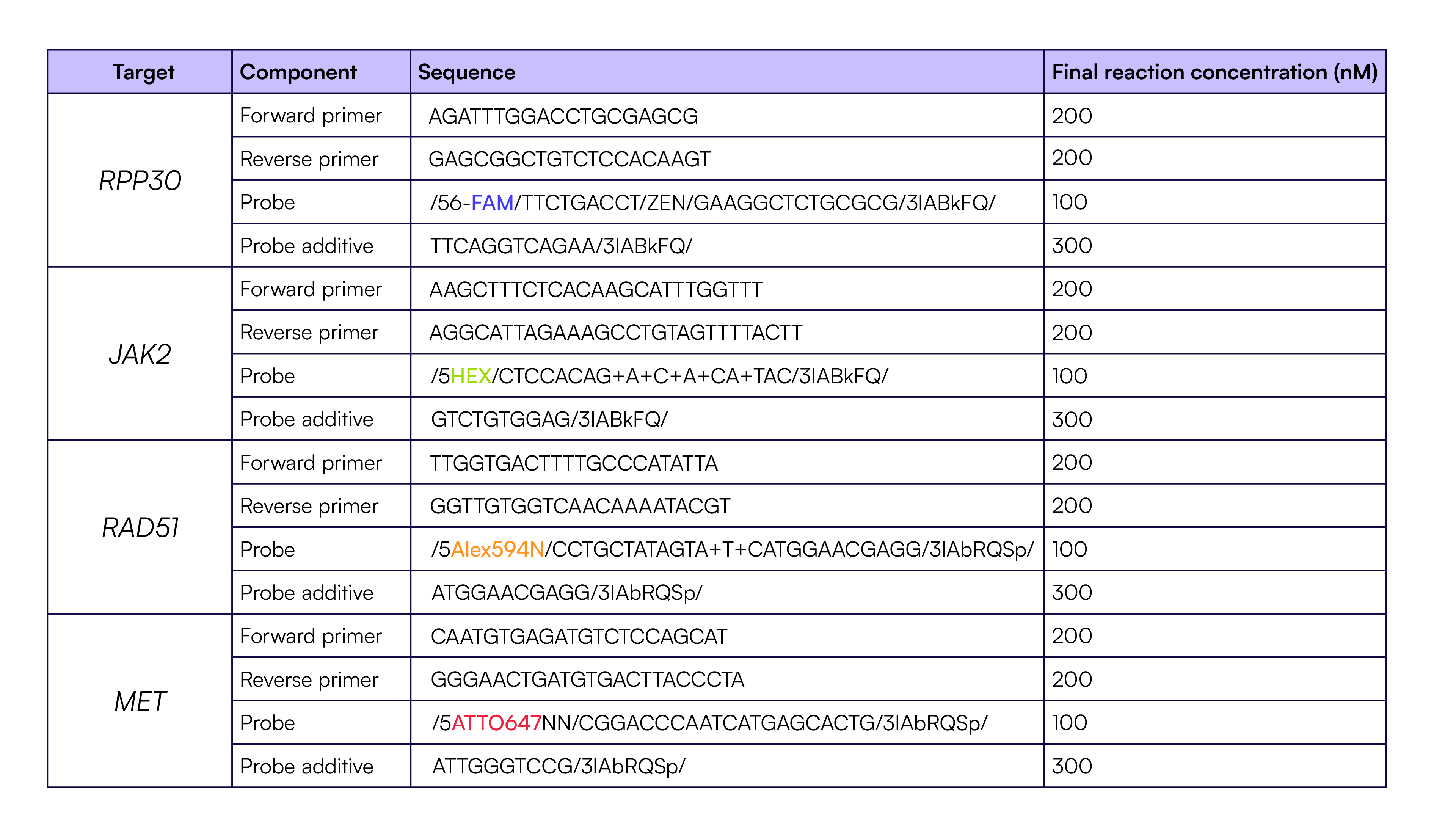
Table 2. Thermal Cycling protocol for qPCR,dPCR, and Countable PCR
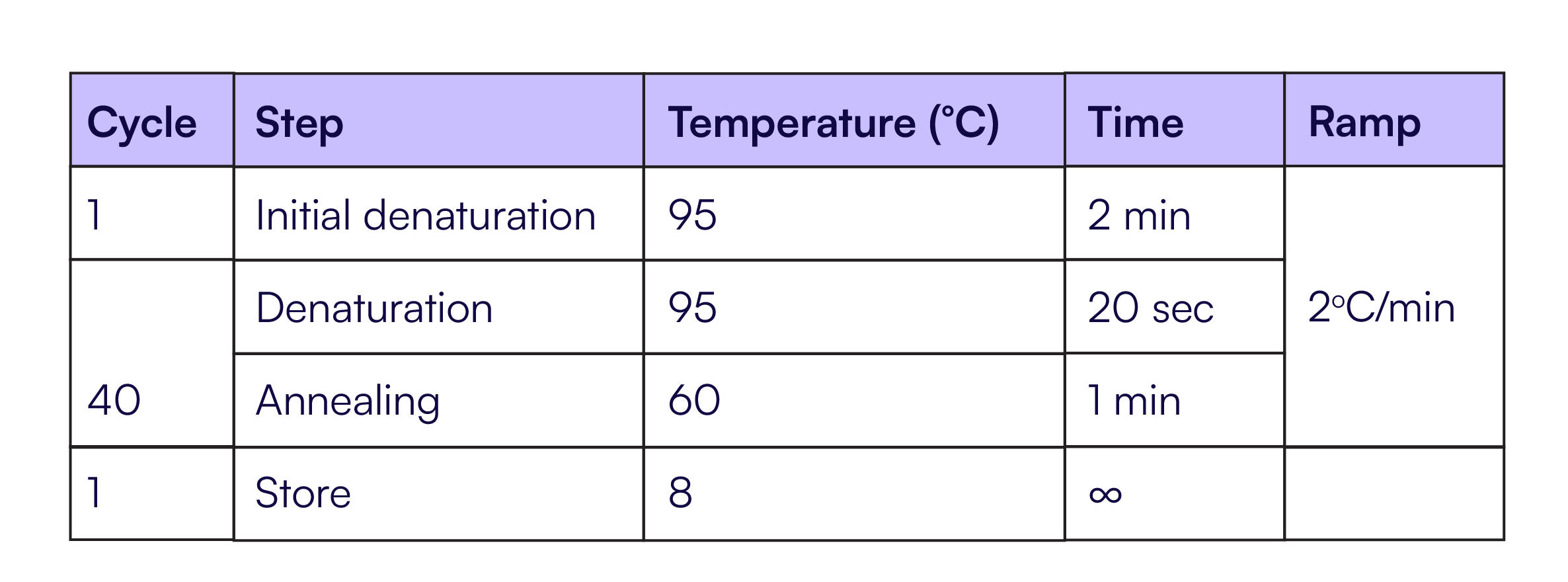
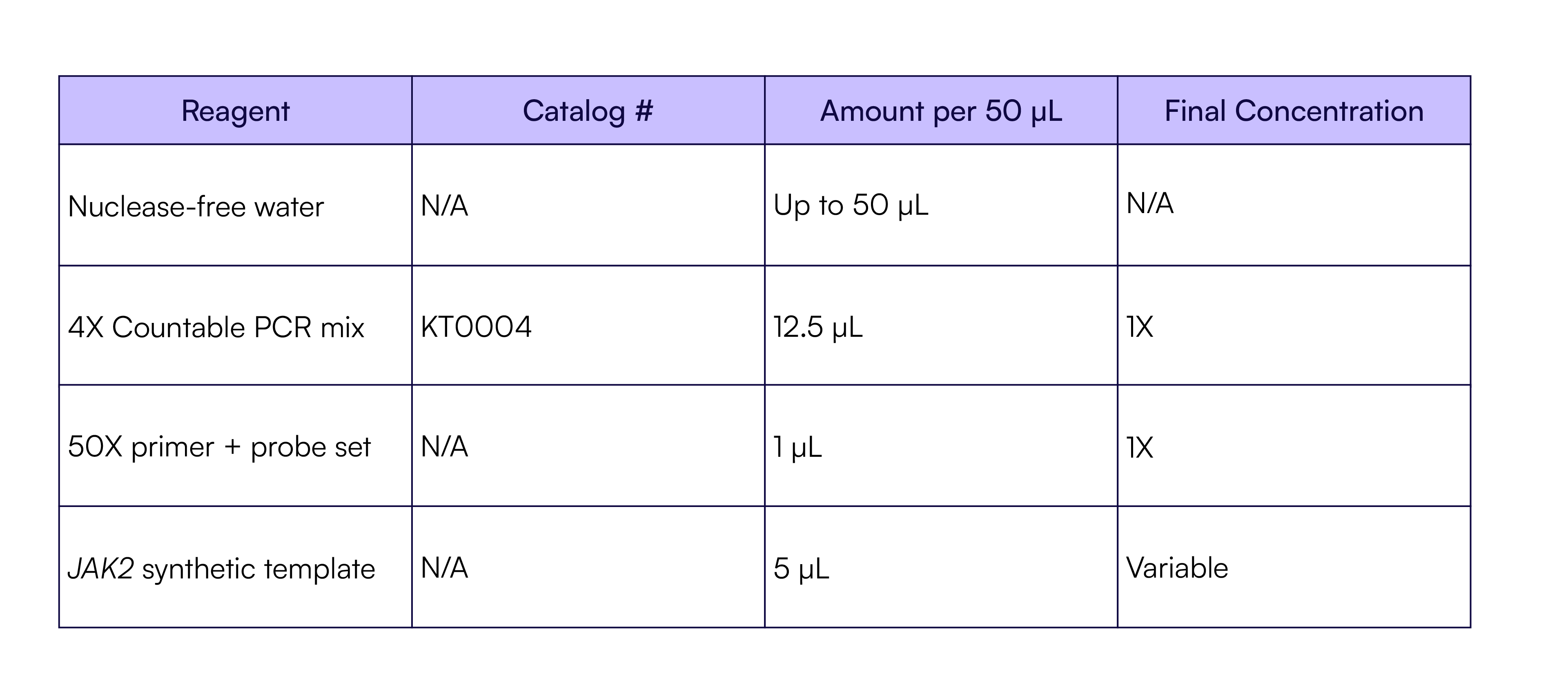
Table 3. RPP30 Countable PCR reaction setup
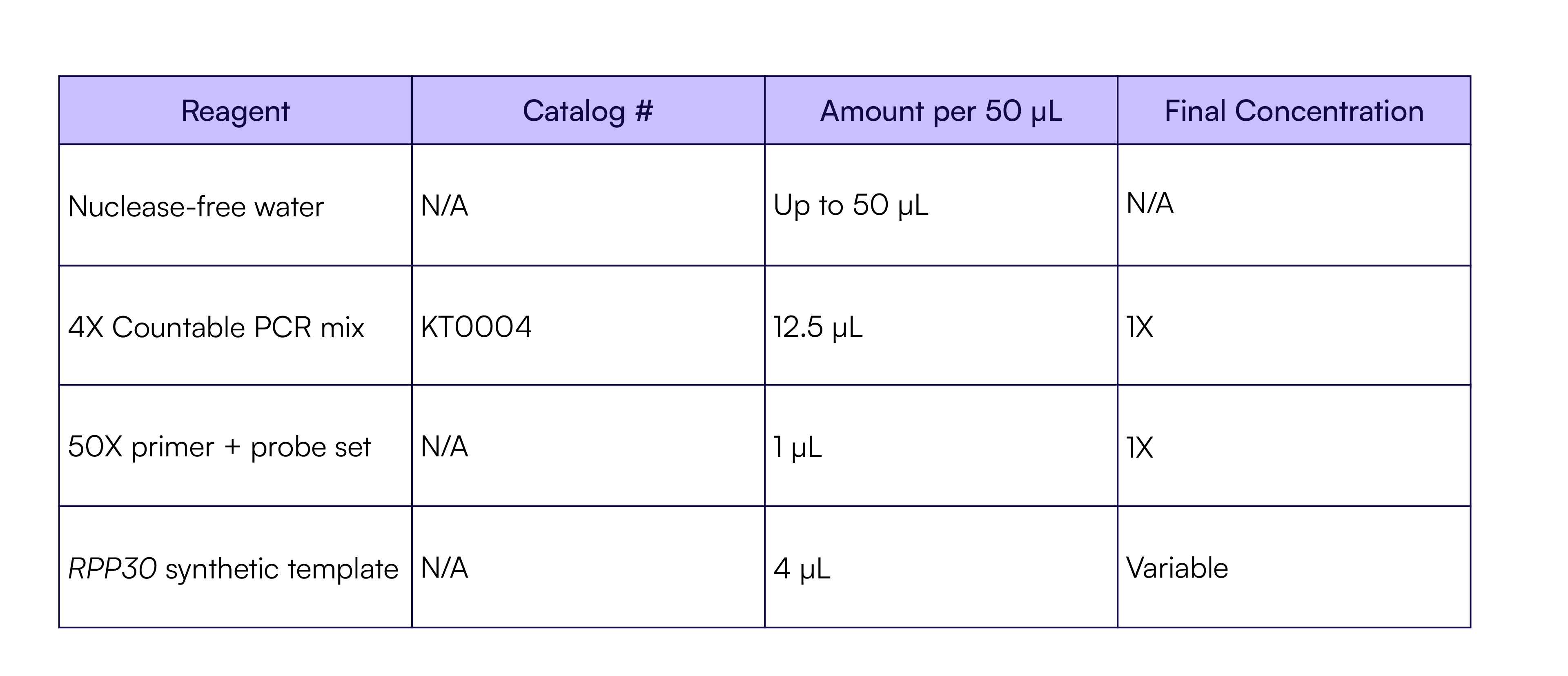
Table 4. Four-plex Countable PCR reaction setup
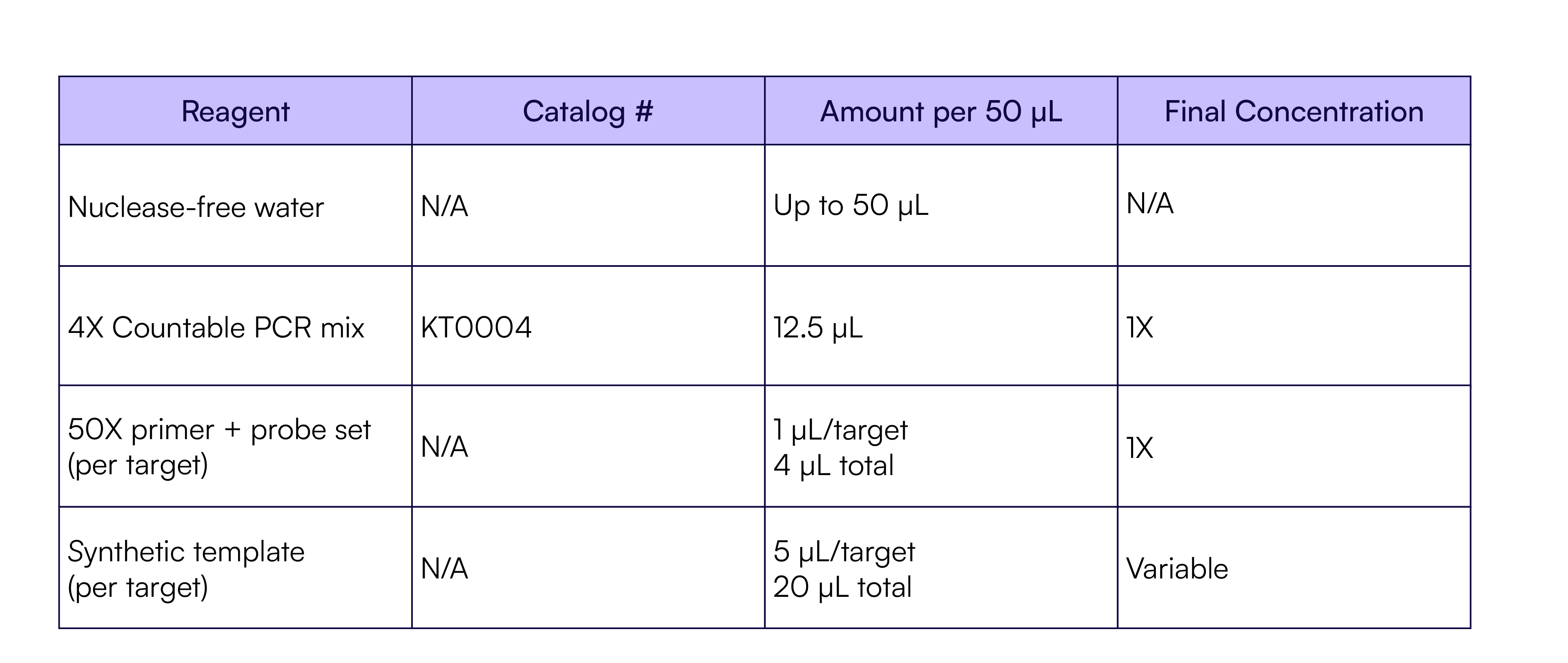
Table 5. JAK2 Countable PCR reaction setup
Results
Countable PCR has a six-log dynamic range, outperforming legacy dPCR systems
Countable PCR’s partitioning matrix creates 30 million compartments, isolating single target molecules and enabling direct counting across six orders of magnitude. This single-molecule partitioning eliminates the need for Poisson-based estimates of single-molecule counts from double and triple-occupied partitions, improving measurement accuracy even at high target counts. Additionally, light sheet imaging allows the Countable Imager to process a larger number of counts within a practical timeframe.
In contrast, dPCR systems have a limited dynamic range due to their reliance on Poisson correction. This limitation is twofold. First, Poisson correction requires a mix of positive (full) and negative (empty) partitions. At high target concentrations, it becomes difficult to maintain enough negative partitions to apply the correction accurately. Second, many dPCR systems are constrained by the number of partitions they generate and process within a practical experiment time.
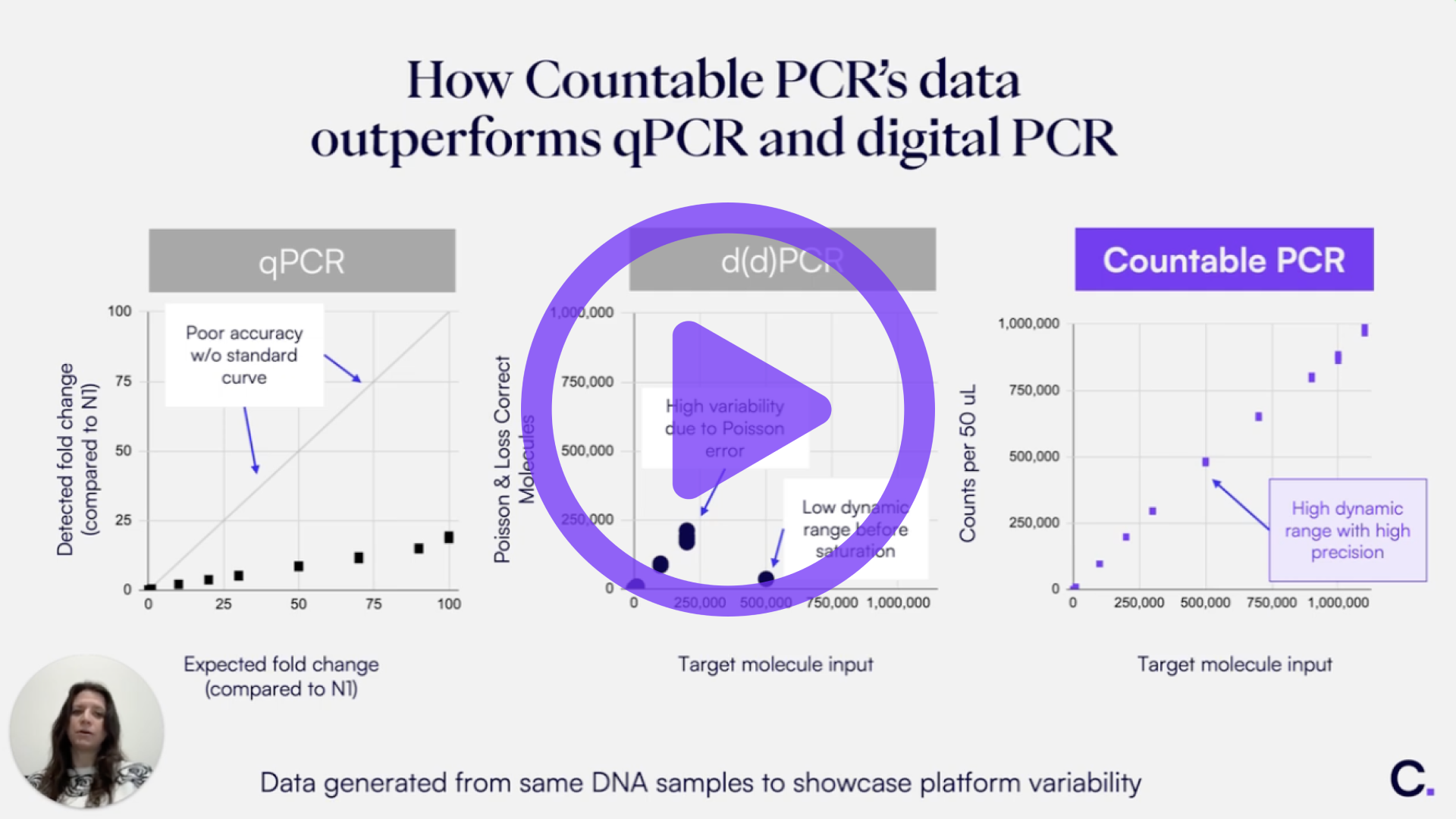
We compared the reported counts from nanoplate dPCR, droplet dPCR, and Countable PCR systems (Figure 1). While all three instruments demonstrated good linearity, only Countable PCR quantified up to 1 million molecules. In contrast, the nanoplate and droplet dPCR systems were limited to approximately 300,000 and 200,000 counts, respectively. With only about 17,000 partitions in the droplet and 25,000 in the nanoplate dPCR system, these legacy dPCR systems quickly saturate as the number of positive partitions approaches their limits. The droplet dPCR system erroneously reports 70 million counts when exceeding its dynamic range, while the nanoplate dPCR system reports 0 counts under the same conditions.

Countable PCR enables enhanced multiplexing capability through a six-log dynamic range
Beyond its six-log dynamic range, Countable PCR’s partitioning matrix enables enhanced multiplexing. By ensuring each partition contains only a single target molecule, it minimizes amplification bias that often occurs for multi-template partitions. Additionally, single-molecule encapsulation simplifies the interpretation of multiplexed reactions, as each partition definitively contains only one target within the 3D matrix.
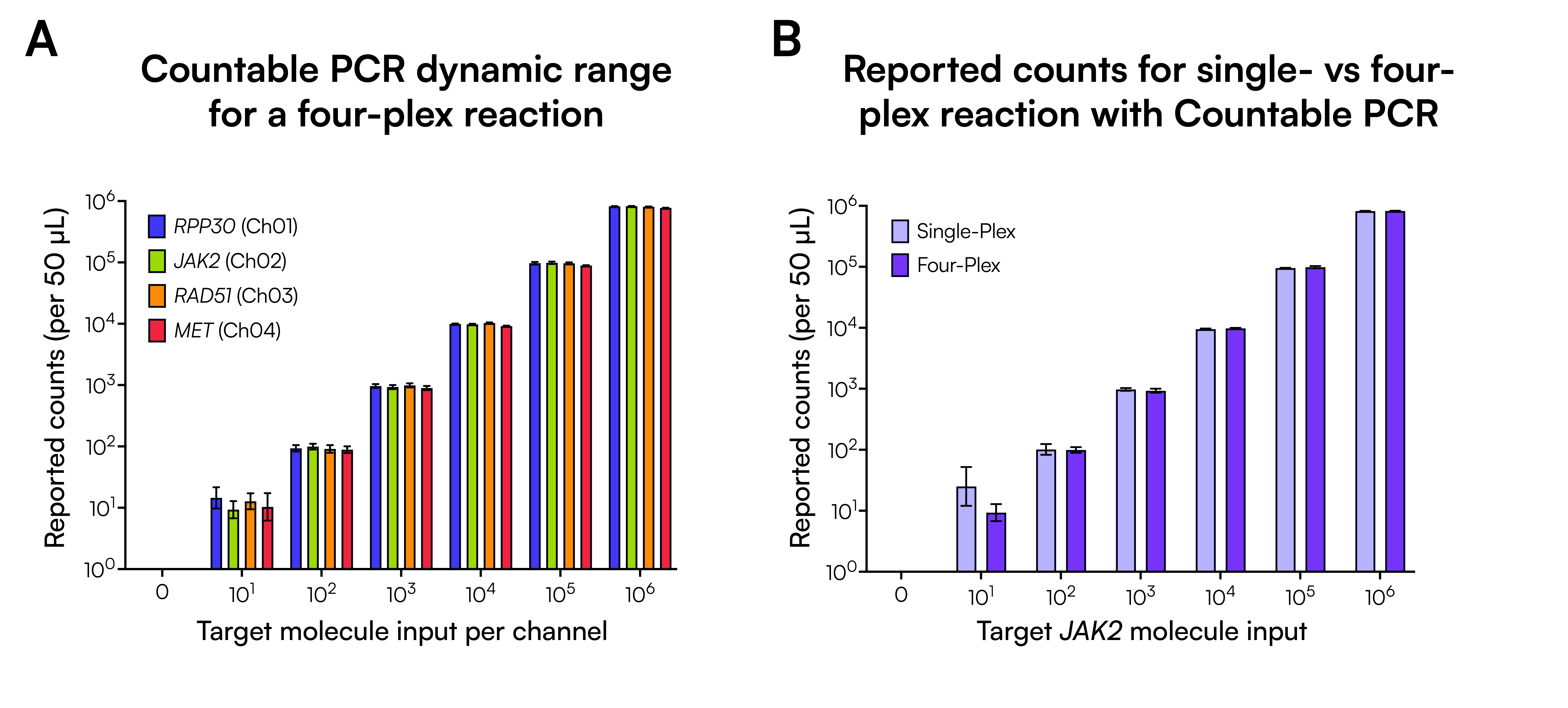
We performed Countable PCR on a series of four-plex reactions containing pooled double-stranded DNA targets RPP30, JAK2, RAD51, and MET, corresponding to Ch01, Ch02, Ch03, and Ch04, respectively (Figure 2A). The pooled templates were serially diluted to produce final counts for each channel ranging from 10 to 1 million. All four channels reported a high degree of linearity (R2 > 0.9995) over a six-log dynamic range.
To show that the four-plex design does not compromise single-plex performance, we ran single-plex JAK2 reactions using the same diluted template (Figure 2B). Across the range of 100 to 1 million counts, the average difference between single-plex and four-plex counts was minimal (2.9%), confirming that Countable PCR accurately quantifies multiple targets in a single reaction without affecting individual channel results.
Conclusion
Countable PCR delivers a broader dynamic range and a robust multiplexing capability. Here we demonstrate:
- Single-molecule direct quantification across a six-log dynamic range without the need for Poisson correction.
- Multiplexing across the full dynamic range without the need to balance amplification biases between targets.
Together, these results highlight Countable PCR as a robust platform for direct quantification of multiple targets over a much broader dynamic range than conventional dPCR-based methods.This enables simpler and more accurate comparison of multiple targets within a single reaction, particularly valuable for rare event detection.
%20option%202.png)


.png)