A single-molecule PCR framework bridging low-plex PCR and NGS for biomarker validation at 16-plex and beyond
Abstract
Next-generation sequencing (NGS) has enabled the discovery of thousands of candidate biomarkers across a wide range of diseases and traits. Translating these discoveries into practical, quantitative assays remains a major challenge. Conventional qPCR and dPCR platforms are widely used for low-plex validation, but their limited multiplexing capacity constrains scalability. Moreover, high-plex PCR assays typically require extensive optimization to balance amplification efficiencies and minimize cross-reactivity, limiting their utility for large-scale validation and clinical assay development. A quantitative platform that combines high multiplexing, digital precision, and efficient use of limited samples is needed to close this gap.
We describe a single-molecule quantitative PCR approach, Countable PCR, designed to address these challenges. The method isolates and amplifies up to one million individual DNA molecules in parallel without sample loss, followed by high-resolution 3D light-sheet imaging for digital counting. Operating in the single-molecule regime enables precise quantification and straightforward multiplexing, as each molecule amplifies independently without inter-target competition. Using ten spectrally distinct fluorophores, this system detects ten independent targets simultaneously on a four-color imaging platform. By applying combinatorial fluorophore labeling, multiplexing is extended beyond twenty targets within a single reaction.
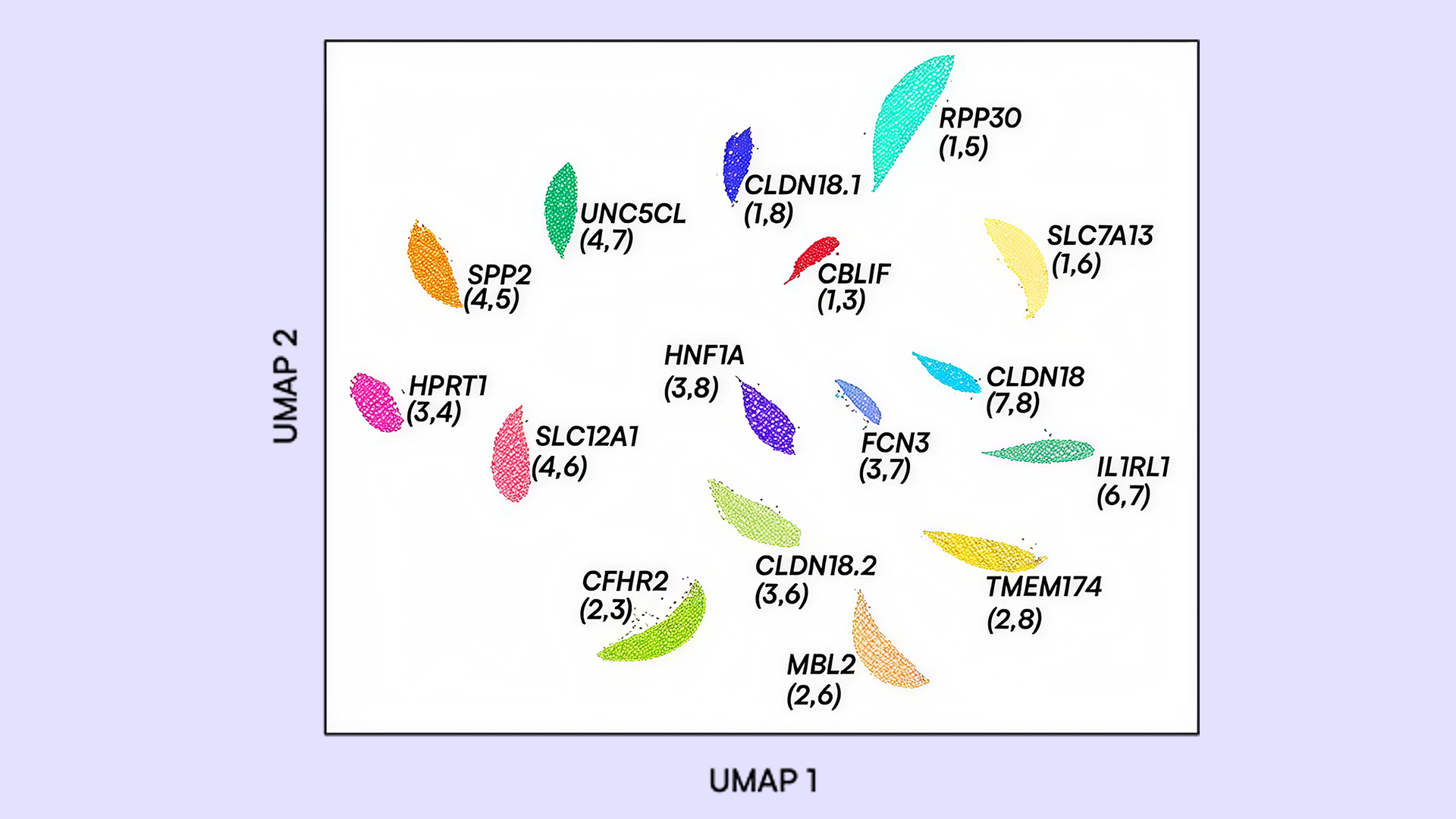
We demonstrate applications including a >20-plex respiratory pathogen detection panel and a 16-plex gene expression panel, both developed with minimal optimization. Quantitative results show concordance with single-plex assays and maintain high target specificity. Data from these panels were further analyzed using multivariate and high-dimensional visualization methods such as UMAP, analogous to approaches used in single-cell transcriptomic analysis, enabling intuitive interpretation of multiplexed results.
This single-molecule PCR framework establishes a scalable strategy for high-plex quantitative analysis with NGS-comparable precision. It provides a generalizable approach for accelerating biomarker validation and supports the development of multi-gene assays for cancer subtyping, prognostic scoring, pathogen detection, and other applications requiring rapid, high-plex quantification.

