Multiplex Detection of BCR-ABL1 Fusion Transcripts in a Streamlined Single-Tube Workflow Using PCR-Based Single Molecule Counting
Multiplex Detection of BCR-ABL1 Fusion Transcripts in a Streamlined Single-Tube Workflow Using PCR-Based Single Molecule Counting

Abstract
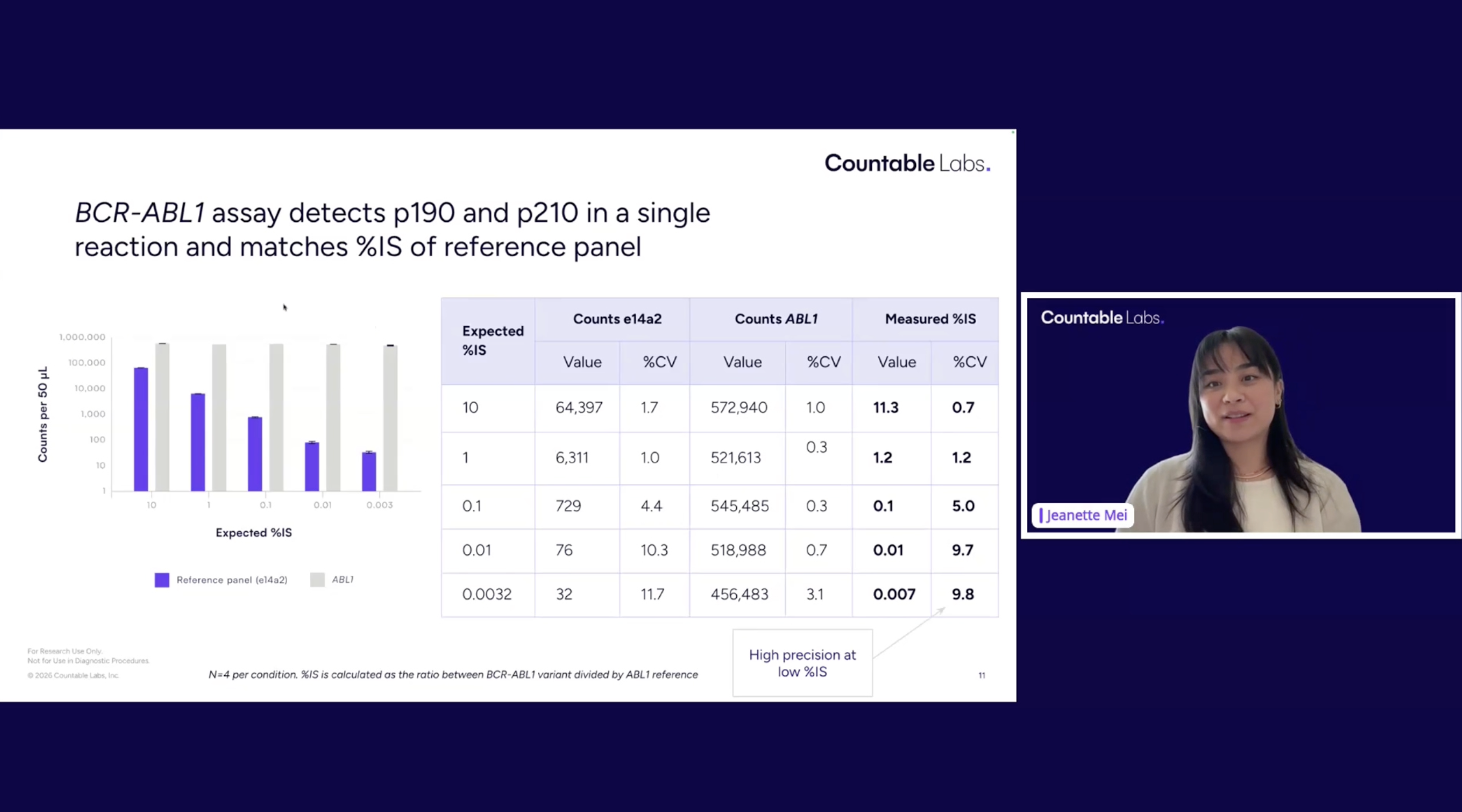
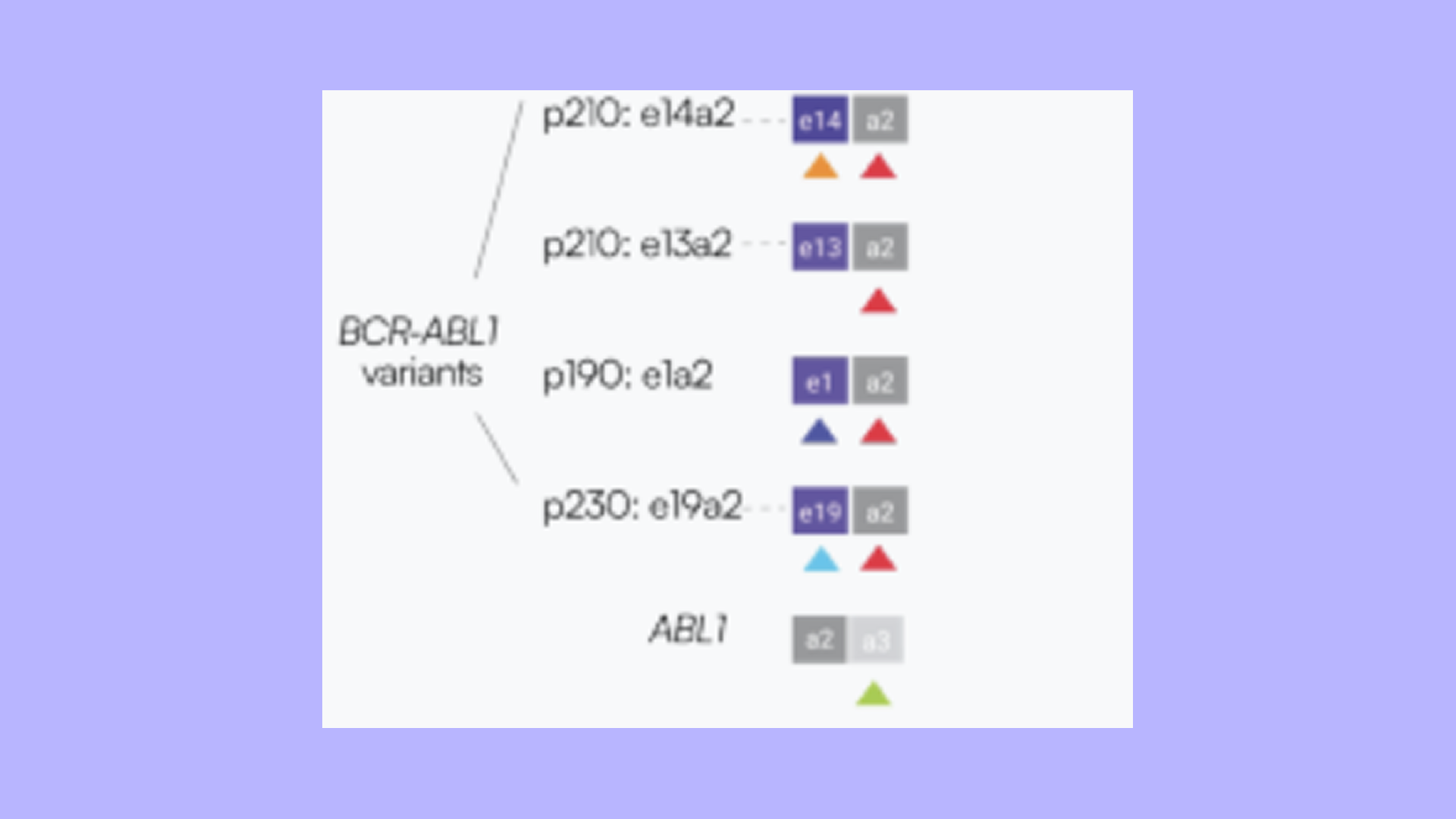
Accurate quantification of BCR-ABL1 fusion transcripts is essential for monitoring minimal residual disease (MRD) in chronic myeloid leukemia (CML), where distinguishing between isoforms (p210, p230, p190) is clinically important for diagnosis, prognosis, and treatment monitoring. Traditional detection methods rely on two-step workflows involving reverse transcription (RT) followed by PCR amplification, either via qPCR or digital PCR (dPCR). While dPCR enables absolute quantification without standard curves, its limited dynamic range and multi-step workflow present challenges for streamlined isoform detection. We developed a one-step multiplex RNA assay for the direct detection and molecular counting of p210, p230, and p190 isoforms, even amid abundant wild-type ABL1 molecules, within a single tube. The entire workflow, including RT-PCR and counting, occurs without sample transfers, reducing hands-on time and contamination risk. Using 3D light-sheet microscopy, molecules are directly counted by imaging the complete 50 µL reaction volume, eliminating dead volume and partitioning statistics. When BCR-ABL1 RNA containing each isoform was serially diluted into wild-type ABL1 background at varying ratios, wild-type ABL1 counts remained stable, near one million molecules, while BCR-ABL1 counts decreased proportionally with dilution.This highlights the highly sensitive and specific detection of RNA isoforms across a 6-log dynamic range with low variance (<10% CV), enabling monitoring of both abundant and rare transcripts within a single tube. This single-tube approach provides a streamlined alternative for sensitive detection of all BCR-ABL1 isoforms in one multiplexed reaction, with potential applications in clinical assessment and therapeutic decision-making in CML.